Table of Contents
10x Genomics Visium Submission
10X Visium samples are processed across both the Histopathology and Genomics cores, so prior to submission, please do contact Single Cell Team Histology
We would recommend arranging to discuss your experimental aims with both teams by discussing this at a experimental design meeting (EDM): Request EDM Here
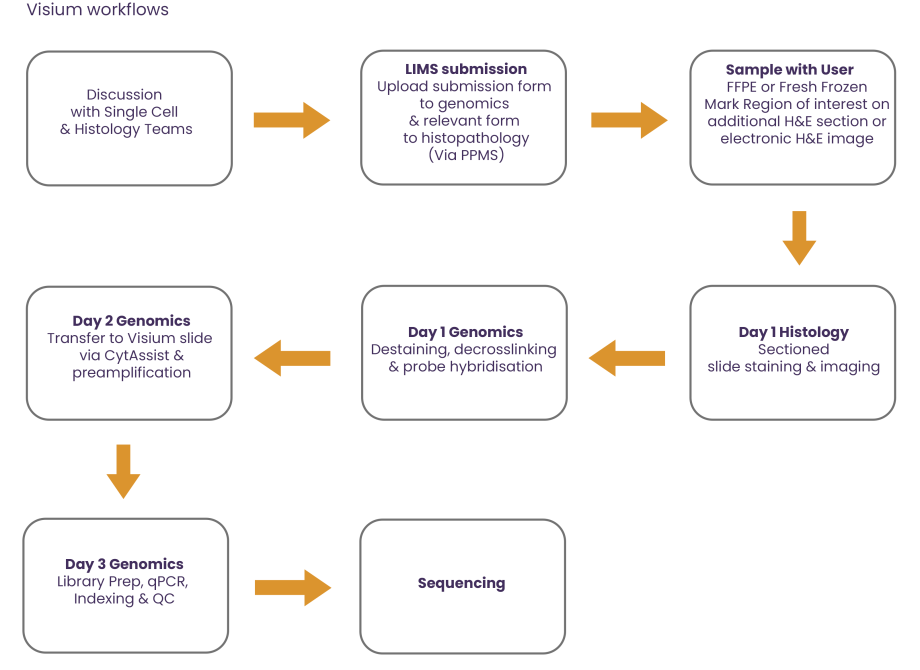
Overview of the Visium & Visium HD workflows
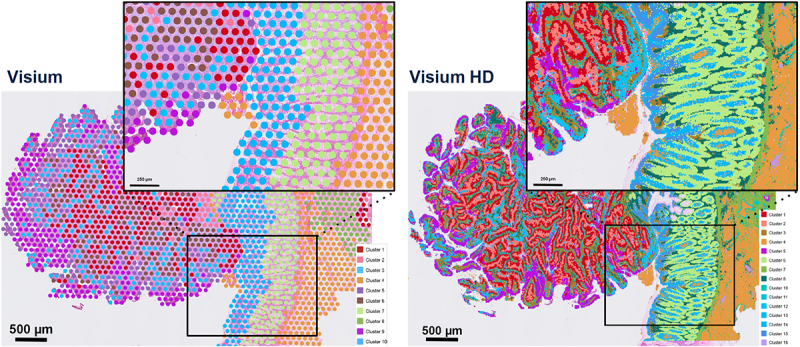
Visium provides a whole transcriptome snapshot of gene expression across a tissue section, allowing you to identify spatial patterns of gene expression within different cell types and tissue regions. With Visium HD a significantly higher resolution is achievable essentially providing “single-cell scale” analysis by capturing gene expression data within very small, densely packed regions of a tissue section when compared to the larger spatial bins of “standard” (SD) Visium (1-10cells).
Visium options available
| Workflow | Kit Used | Slide used for submission | Method of mRNA capture | Spot-based array |
|---|---|---|---|---|
| Visium HD (FFPE) | Visium HD slide & Cassette, Visium HD reagent kit, Visium Human or mouse probe kit | 6.5 x6.5mm section fixed on a regular superfrost slide | Whole transcriptome probe capture (Human & Mouse probe kits available) | lawn of 2um, recommended bin size for analysis 8uM, single cell resolution |
| Visium HD (fresh frozen) | Visium HD Slide & Cassette, Visium HD reagent kit, Visium Human or Mouse Probe Kit | 6.5 x6.5mm section fixed on a regular superfrost slide | Whole Transcriptome probe capture (Human & Mouse probe kits available) | lawn of 2um, recommended bin size for analysis 8uM, single cell resolution |
| Visium CytAssist (FFPE) | Visium Slide & cassette, FFPE Reagent Kit, Probe Kit | FFPE tissue section of either 11 x 11 mm or 6.5 x 6.5mm in area, fixed onto regular superfrost slide | Whole Transcriptome probe capture (Human & Mouse probe kits available) | Spot size 55um 11 x 11mm 14000 spots 6.5mm x 6.5mm 5000 spots, average 1-10 cells resolution |
| Visium CytAssist (Fresh Frozen) | Visium Slide & cassette, FFPE Reagent Kit, Probe Kit | Fresh tissue section of either 11 x 11mm or 6.5 x 6.5 mm in area, fixed onto regular superfrost slide | Whole Transcriptome probe capture (Human & Mouse probe kits available) | Spot size 55um 11 x 11mm 14000 spots 6.5mm x 6.5mm 5000 spots, average 1-10 cells resolution |
| Visium Spatial Gene Expression Manual (FFPE) | Visium Spatial Gene Expression Slide Kit, Visium FFPE Reagent Kit, Probe Kit | 4 capture areas available for tissue sections 6.5 x 6.5mm in area, fixed onto a spatial gene expression slide (provided by 10X) | Whole Transcriptome probe capture (Human & Mouse probe kits available) | Spot size: each capture area has ~5000 spots, average 1-10 cells resolution |
| Visium Spatial Gene Expression Manual (Fresh Frozen) | Visium Spatial Gene Expression Slide & Kit, Probe Kit | 4 capture areas available for tissue sections 6.5 x 6.5mm in area, fixed onto a spatial gene expression slide (provided by 10X) | Poly(A) tail capture | Spot size: each capture area has ~5000 spots, average 1-10 cells resolution |
| Visium CytAssist Spatial Gene & Protein Expression | Visium Slide & Cassette, FFPE Reagent Kit, Probe Kit, Immune Cell Profiling Panel, Add on antibodies (if applicable), Visium Protein Core Reagents (CITE-Seq) | FFPE tissue section of either 6.5 x 6.5mm in area, fixed onto regular superfrost slide | Whole Transciptome probe capture (Human & Mouse probe kits available) | Spot size: 11 x 11mm 14000 spots 6.5mm x 6.5mm 5000 spots, average 1-10 cells resolution |
Make sure you provide additional H&E section or electronic version of H&E image with marked area of interest (ROI) according to the selected workflow (11 x 11mm or 6.5 x 6.5mm) to guide placement of the gasket and the alignment of the superfrost slide against the Visium slide capture array.
How to Complete the 10x Visium Submission Form
Here is the guide how to fill it out. The first few fields are related to which workflow you want your samples to go through.
- Workflow: This tells us if you are using Fresh Frozen or FFPE tissue, if you would like to use Visium HD or standard Visium with CytAssist or Visium manual method, and what size of tissue section you are submitting (either 11 x 11mm or 6.5 x 6.5mm).
- Ready to sequence:
- Yes: if you have submitted all your current samples and are ready to pool and sequence. Please contact us and let us know which samples you want pooled. Please email the single cell email address (singlecell@cruk.cam.ac.uk) with the sequencing information (sequencer, flowcell type, number of lanes, reads per spot).
- No: if you are submitting multiple samples over a time period. You do not need to let us know the sequencing information at this point.
- Probe: Please specify either human or mouse whole transcriptome probes accordingly to the species you work with. Leave this blank if your probe is not in the list of options
- Sample source: Fresh, Fresh Frozen, FFPE or Other.
- HTA: This only applies if the tissue falls under the human tissue act. If yes, please contact the single cell team as we need additional information before accepting this type of sample.
- Species: Choose from the drop down menu. Leave this blank if your probe is not in the list of options.
- Billing Information: This is critical for correct invoicing: please follow the popup instructions in the field. If you do not have a SWAG/SWAK code, please contact us to discuss.
- PO number: Only enter a PO number if applicable to your department or organisation. Leave PO number blank if it does not apply! Resist the temptation to put “N/A” or “None”: it will complicate your bills.
Individual Sample Information
- Name: Name of your sample.
- RGT Number: This ID is related to human tissue and is generated when sample is processed by Tissue Bank or being submitted to CRUK CI (using Achiever Medical). Format of RGT00xxxxxx.
- RGT Aliquot Number: This provided by histopathology or Tissue Bank (if they generate aliquots of original tissue). Format of three-digit code after the RGT number: RGT00xxxxxx-001 (leave empty if you do not have that information).
- Slide ID: Any additional slide ID provided eg. CRU number or what is written on the slide.
- Capture Area (%): If it is known what percentage of the 6.5 x 6.5mm or 11 x 11mm capture area your tissue covers, please write this here (leave empty if you do not have that information).
- DV200: Please provide this if you have done an RNA extraction and TapeStation QC on a tissue section from the same block, to give us an idea of the RNA quality of your tissue sample (leave empty if you do not have that information).
- Species: only complete this if it is different from the species provided in the sample set information.
- Reads per spot: How many sequencing reads do you want per spot of the Visium slide? – this indicates the depth of sequencing you would like.
For Visium HD, 10x Genomics recommend 275M reads per capture region. For standard Visium with CytAssist or manual, 10x Genomics suggest 25,000 reads per spot for FFPE tissue, and 50,000 reads per spot for fresh frozen on their guidelines but it is up to you! How many reads per spot you want does influence what flow cell you will need to use for sequencing, so if you would like some guidance, please do contact the Single Cell team who will be able to advise you.
10X Genomics sequencing configuration recommendations
| Visium Option | Feature | Configuration: Read1/i7/i5/Read2 | Sequencing Depth |
|---|---|---|---|
| Spatial Gene Expression | Fresh Frozen Direct Placement v1 | 28/10/10/90 | 50K read pairs / spot |
| FFPE Direct Placement v1 | 28/10/10/50 | 25K read pairs / spot | |
| CytAssist Gene Expression FF,FFPE,FxF v2 | 5K read pairs / spot | ||
| CytAssist Gene & Protein Expression | 25K read pairs / spot 5K Protein |
||
| HD Spatial Gene Expression | HD Gene Expression | 43/10/10/50 | (275M read pairs) x (% Capture Area covered by tissue) |
- Submission comments: Anything that needs to be communicated and might be relevant for the experiment like suggestions about pooling before sequencing, type of antibodies used for staining (Spatial gene and protein expression), information about possible low RNA content etc.